Science
Oncogenic Pathways and Iron Phenotypes in Breast Cancer
By Viktor Belay, BS Physics and Mathematics; Sergio Guerra Abril, BS Mathematics and BA Dance; Jacqueline Adams, BS Mathematics; Linoy Kotler, BS Physics and Mathematics; Julia Chifman, Assistant Professor in the American University Department of Mathematics and Statistics
Iron is a ubiquitous element. It is pervasive throughout nature and is critical to almost all life on Earth; even so, this essential element can be toxic due to its ability to exist in multiple oxidation states. In particular, iron has the capability to donate and accept electrons, which is an extremely useful property for many biological processes. At the same time, this activity can facilitate the formation of highly reactive particles known as free radicals. These radicals react with intra- and inter-cellular machinery, often generating undesired biochemical effects. Generally, there is a balance between the production of these radicals and the production of antioxidant enzymes that neutralize free radicals. However, large scale free radical formation may lead to DNA damage and genetic mutations.
To avoid iron toxicity, yet to deliver adequate amounts of iron for biological processes, life on Earth has evolved to have sophisticated machinery that regulates iron at both the systemic and the cellular levels. This machinery is known as iron metabolism. When iron metabolism ceases to function properly, the resulting iron excess or deficit can lead to serious faults in other cellular structures and regulatory systems. These faults are well studied and are known to cause dangerous medical conditions such as neurodegeneration, heart failure, iron-deficiency anemia, and even cancer.
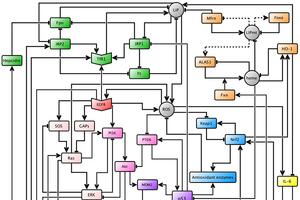
Cellular iron levels and iron associated proteins are differentially expressed in many cancers, and recently, it was reported that high expression levels of iron importer and low levels of iron exporter were associated with a poor prognosis in breast cancer patients. Thus, Julia Chifman’s research aims to understand the cause of iron dysregulation in breast cancer through the study of certain cellular signaling pathways that affect iron homeostasis. Cellular signaling pathways are networks of genes, proteins, enzymes, cytokines, and other cellular species that “communicate” in order to govern the basic functions of cells. Assistant Professor of Mathematics and Statistics Julia Chifman and her collaborators built a predictive mathematical model that connects four key cytosolic cellular signaling pathways in breast epithelial cells: the oncogenic pathway, the oxidative stress response pathway, the iron utilization pathway, and the iron homeostasis pathway. The current network is shown in figure 1.
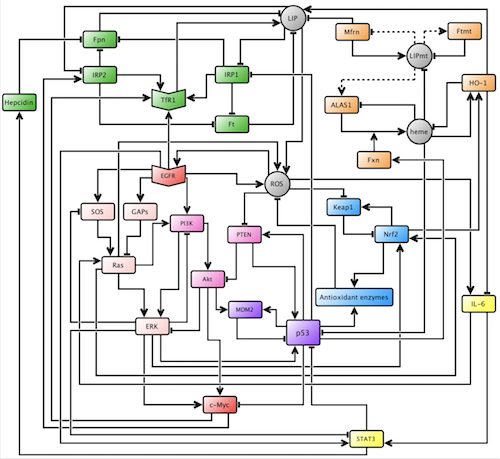
Figure 1.
The model uses two basic operations between species to simulate their responses to each other: activation and inhibition, represented by an arrow and a hammerhead arrow, respectively. Therefore, if one species activates another, a high concentration or activity level will induce an increase in the levels of the second species. On the other hand, if a species inhibits another, a high concentration or activity level will induce a decrease in the levels of the second species. Each of these interactions is encoded through mathematical logic, and the behavior of each node in the network is then expressed through a logical update rule, which is then translated into a polynomial. Once all the polynomials are encoded, one can simulate the network with a computer given initial conditions. In the context of these simulations, initial conditions are certain expression levels of some species that one might want to investigate.
Led by Chifman, our research team worked to better understand and develop the oncogenic pathway portion of the established pathway model. The oncogenic signaling pathway includes cellular species that regulate the growth, differentiation, and survival of cells. Faults in this pathway can lead to downstream alterations in a multitude of cellular systems; namely, defects in the oncogenic pathway are noted to induce modifications in cellular iron homeostasis. Through simulations, Chifman and her collaborators found that overexpression of Ras, a central species in the oncogenic pathway, led to a cancer phenotype in the iron homeostasis pathway.
In the schema of our project, our team sought two central goals: to increase the biological accuracy of the model through the addition of cellular species and intra-pathway connections to the oncogenic pathway, and to conduct new simulations using the updated oncogenic pathway.
In order to determine which cellular species and connections were appropriate to add to the model, we researched biological and biochemical experimental results in scholarly literature. With these findings, we were able to add a total of six new species to the oncogenic pathway and 22 new connections to the model. We added two genes, p53 and the phosphatase tensin homolog (PTEN), which both encode proteins that regulate cell growth and act as tumor suppressors; a transcription factor, the signal transducer and activator of transcription 3 (STAT3), which mediates the expression of genes that are involved in cell growth and apoptosis; a family of enzymes, phosphatidylinositide 3-kinases (PI3K), which are involved in cell proliferation and growth; a protein kinase, protein kinase B (Akt), which is involved in cell apoptosis and glucose metabolism; and a protein, MDM2, which participates in a negative feedback loop with p53. Our contributions are highlighted in figure 2.
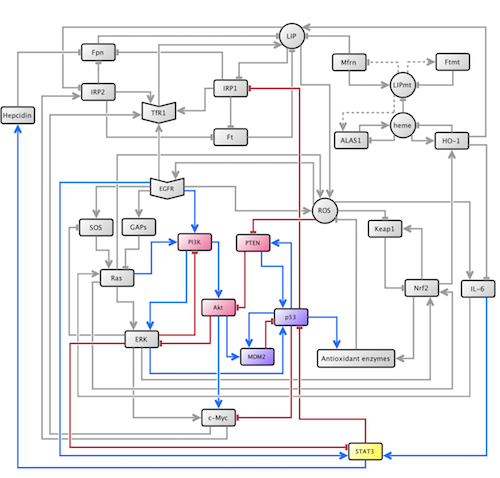
Figure 2.
After constructing polynomials for all of the newly added species, we simulated the model with some particular initial conditions. Our most significant result came with the overexpression of the family of enzymes PI3K; when overexpressed, PI3K not only led to a cancer phenotype in the iron homeostasis pathway, but led precisely to the same phenotype as overexpression of Ras. This result suggests that Ras and PI3K work in a complementary manner in the disruption of iron metabolism in breast cancer.
There are a few main future directions in this project. All of these directions currently aim to make the model more biologically accurate. A significant future step towards higher biological accuracy involves making our model compartmental; that is, we will eventually create a model that considers the effects of cellular signaling between different areas of the cell, such as within mitochondria, on the surface of the cell membrane, in the cytosol, and more. This compartmental model will allow us to track the amount of iron at a specific location or in a particular state by specifying interactions among the compartments, with each compartment having its own network.